Spatially Varying Coefficient Models with Spike-and-Slab Group Lasso
2026 Symposium on Data Science and Statistics, Milwaukee, WI USA
Department of Mathematical and Statistical Sciences
Marquette University
2026-04-29
Why Spatially Varying Coefficients (SVC) Models?
- A single global coefficient may mask or distort spatially heterogeneous relationships.
- SVC models let regression effects change with location: \beta_j(s), not a single \beta_j (Gelfand et al. 2003; A. Stewart Fotheringham, Brunsdon, and Charlton 2002).
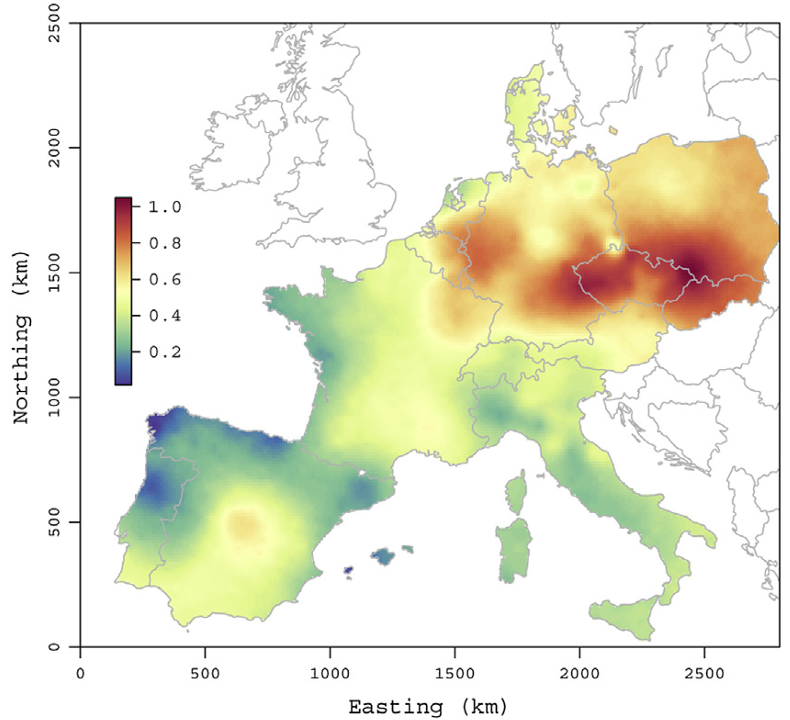
Published SVC applications show this need in geostatistics.
Air pollution (Hamm et al. 2015)
Forestry (Babcock et al. 2015)
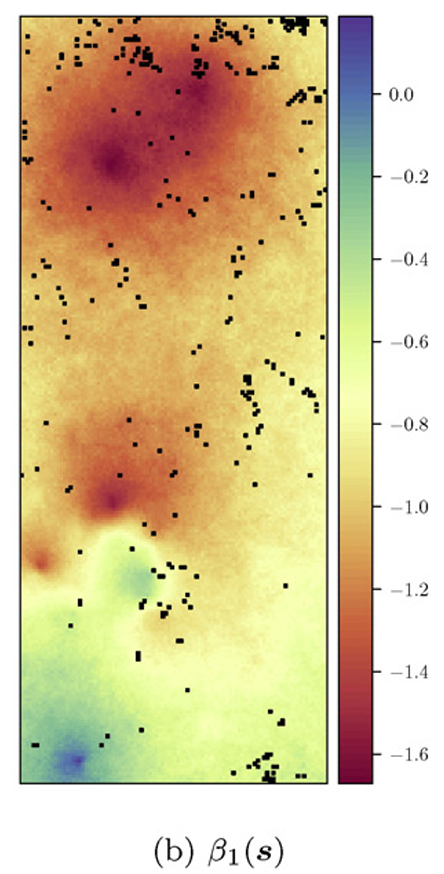
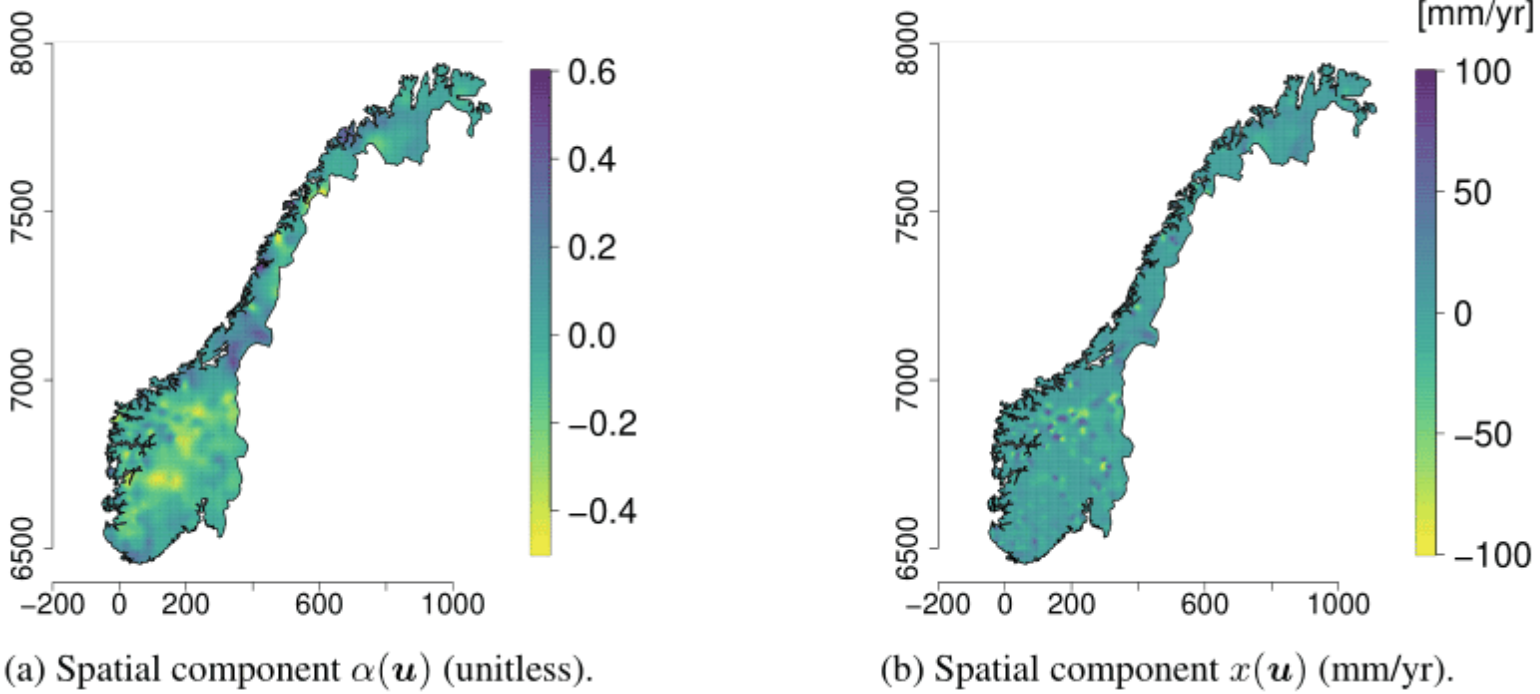
Hydrology (Roksvåg, Steinsland, and Engeland 2022)
Why Selection Matters in SVC Models
Statistical question: which predictors truly need a spatially varying coefficient surface?
- Not every predictor needs a full spatial coefficient surface.
Example. In a county-level diabetes analysis, population density was significant in the model but locally positive in only 0.7% of counties (Hipp and Chalise 2015).
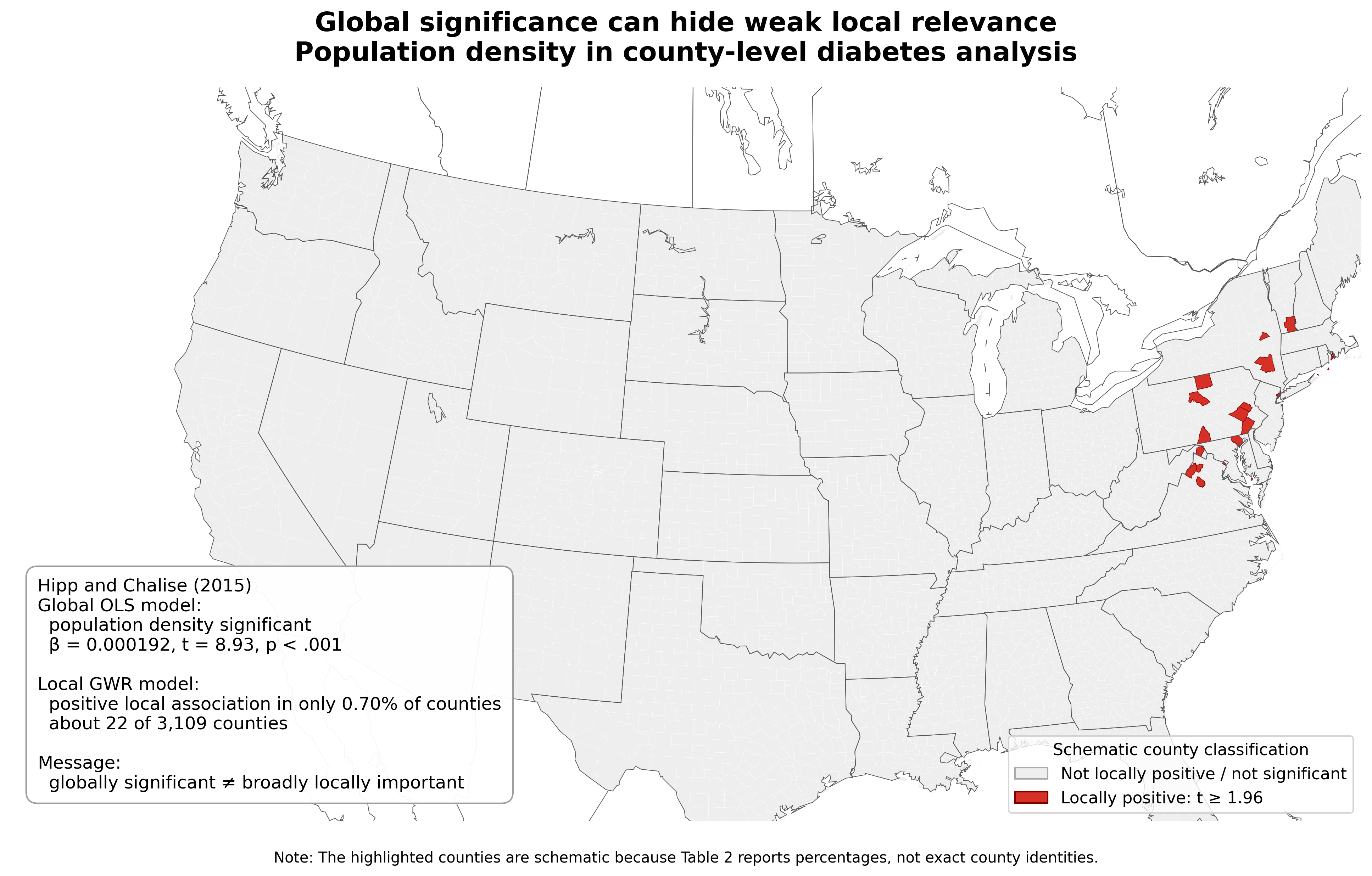
Why Selection Matters in SVC Models
If \beta_j(s) = 0 for all s but x_j is included, but the model tries to estimate a coefficient surface that does not truly exist or scientific meaningful:
-
Estimation (Hastie, Tibshirani, and Friedman (2009); Sestelo et al. (2016))
- Fit random noise instead of real signal
- Spatial smoothing can turn noise into a fake-looking spatial pattern
- Uncertainty in other coefficient surfaces may increase
- Collinearity with useful predictors can make important effects harder to estimate
-
Prediction (Hastie, Tibshirani, and Friedman (2009); Ishwaran and Rao (2005))
- Increase prediction variance
- Reduce out-of-sample predictive accuracy
Why Selection Matters in SVC Models
If \beta_j(s) = 0 for all s but x_j is included, but the model tries to estimate a coefficient surface that does not truly exist or scientific meaningful:
-
Interpretation (Gelfand et al. (2003); Greve and Fischl (2018); Brodoehl et al. (2020))
- Misread as scientific evidence
- Suggest that predictor j matters in some locations when it does not
- Apparent spatial regions of importance may be false positives
-
Computational
- More parameters must be estimated
- Slower MCMC and longer computation
Gap in Existing Methods
Standard selection
✅ sparse predictors
❌ spatially varying effects
Lasso and group lasso select variables or groups, but their standard forms use global coefficients (Tibshirani 1996; Yuan and Lin 2006).
SVC / GWR
✅ spatially varying effects
❌ automatic surface selection
SVC and GWR estimate location-specific or process-based coefficient variation (Gelfand et al. 2003; A. Stewart Fotheringham, Brunsdon, and Charlton 2002).
Penalized spatial methods
✅ shrinkage / local selection
❌ posterior uncertainty as a default output
Geographically weighted lasso is optimization based (Wheeler 2009).
Bayesian SVC selection
✅ model uncertainty
❌ a general scalable surface-selection template
Existing Bayesian SVC selection is often model- or application-specific (Reich et al. 2010; Shang and Clayton 2012; Boehm Vock et al. 2015; Mu et al. 2021).
Need: spatial flexibility + predictor-level surface selection + Bayesian uncertainty quantification.
SSGL-SVC: Our Solution
- Represent each spatially varying coefficient surface with a low-rank basis expansion
\beta_j(s) = \sum_{h=1}^H B_h(s)\alpha_{jh}, \quad j = 1, \dots, p
- Apply spike-and-slab group lasso shrinkage to the basis coefficients for each predictor
\boldsymbol{\alpha}_j = (\alpha_{j1}, \ldots, \alpha_{jH})^\top
-
Selection operates at the predictor-surface level (not local regions)
- keep predictor j if its coefficient surface is supported by the data
- shrink the entire surface toward zero if the predictor is weak
- The result is a scalable Bayesian SVC model with interpretable spatial surfaces and posterior uncertainty
SSGL-SVC: Basis Representation and Low-Rank Design
A separate coefficient at every location for every predictor
Gaussian process dimension grows like p \times n location-specific coefficients
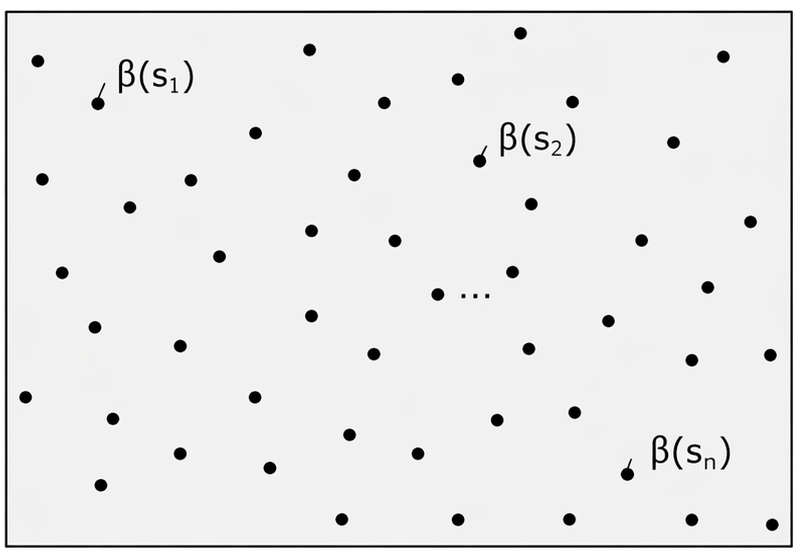
SSGL-SVC: Basis Representation and Low-Rank Design
Approximate each coefficient surface with H basis functions, where H \ll n
Predictor x_j get one coefficient block (\eta_{j1}, \dots, \eta_{jH})
Dimension changes from pn location-level coefficients to pH grouped basis coefficients
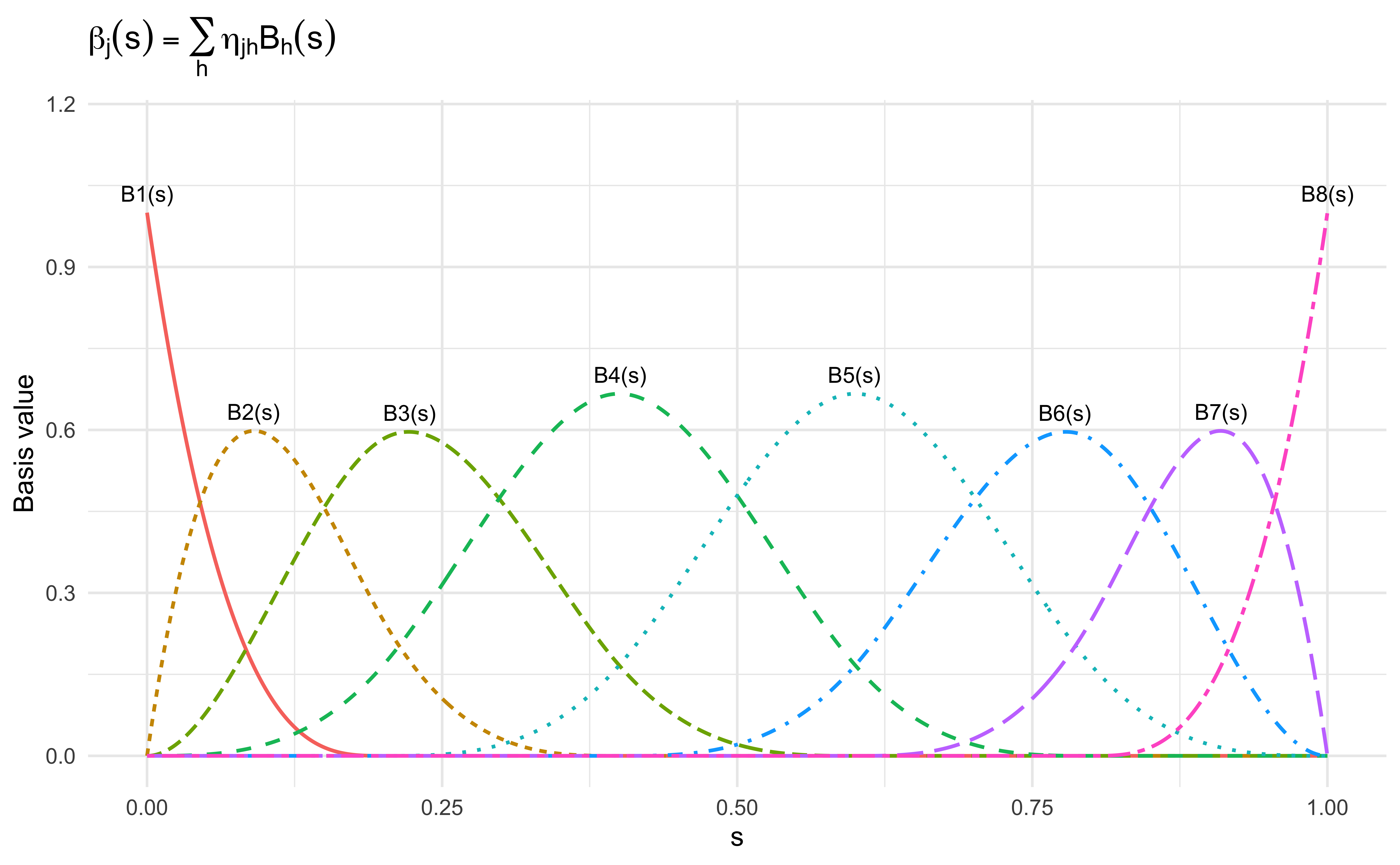
SSGL-SVC: Basis Representation and Low-Rank Design
Turn SVC Into a Grouped Regression
y_i = \alpha_0 + \sum_{j=1}^{p} x_{ij}\,\beta_j(s_i) + \varepsilon_i, \qquad \varepsilon_i \stackrel{iid}{\sim} N(0,\sigma^2), \qquad i = 1, \dots, n
\beta_j(s) = \mathbf{B}(s)^\top \boldsymbol{\eta}_j, \qquad \mathbf{B}(s) = \big(B_1(s),\ldots,B_H(s)\big)^\top
\boldsymbol{\eta} = \big(\boldsymbol{\eta}_1^\top,\ldots,\boldsymbol{\eta}_p^\top\big)^\top, \qquad \mathbf{y} = \alpha_0\mathbf{1}_n + \mathbf{Z}\boldsymbol{\eta} + \boldsymbol{\varepsilon}
\mathbf{Z} = [\mathbf{Z}_1\ \cdots\ \mathbf{Z}_p], \qquad \mathbf{Z}_j = \operatorname{diag}(x_{1j},\ldots,x_{nj})\,\mathbf{\Phi},
where \mathbf{\Phi}_{n\times H} = \left( \mathbf{B}(s_1)^\top, \mathbf{B}(s_2)^\top \dots, \mathbf{B}(s_n)^\top \right)^\top
SSGL-SVC: Basis Representation and Low-Rank Design
\boldsymbol{\eta} = \big(\boldsymbol{\eta}_1^\top,\ldots,\boldsymbol{\eta}_p^\top\big)^\top, \qquad \mathbf{y} = \alpha_0\mathbf{1}_n + \mathbf{Z}\boldsymbol{\eta} + \boldsymbol{\varepsilon}
\mathbf{Z} = [\mathbf{Z}_1\ \cdots\ \mathbf{Z}_p], \qquad \mathbf{Z}_j = \operatorname{diag}(x_{1j},\ldots,x_{nj})\,\mathbf{\Phi},
- Use the same basis matrix \mathbf{\Phi}_{n\times H} for all predictors; multiply by each covariate to form grouped design blocks
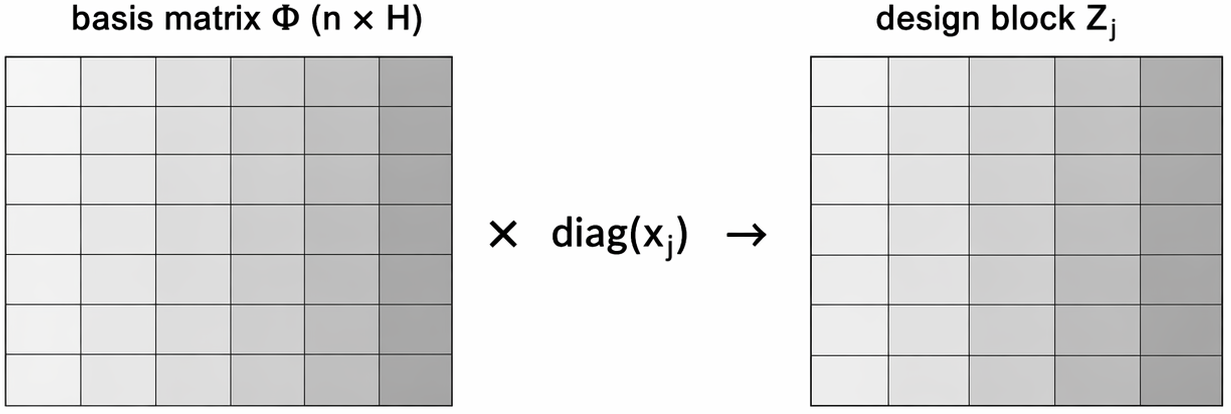
SSGL-SVC: Spike-and-Slab Group Lasso Prior
Basis expansion improves scalability, but does not solve predictor selection.
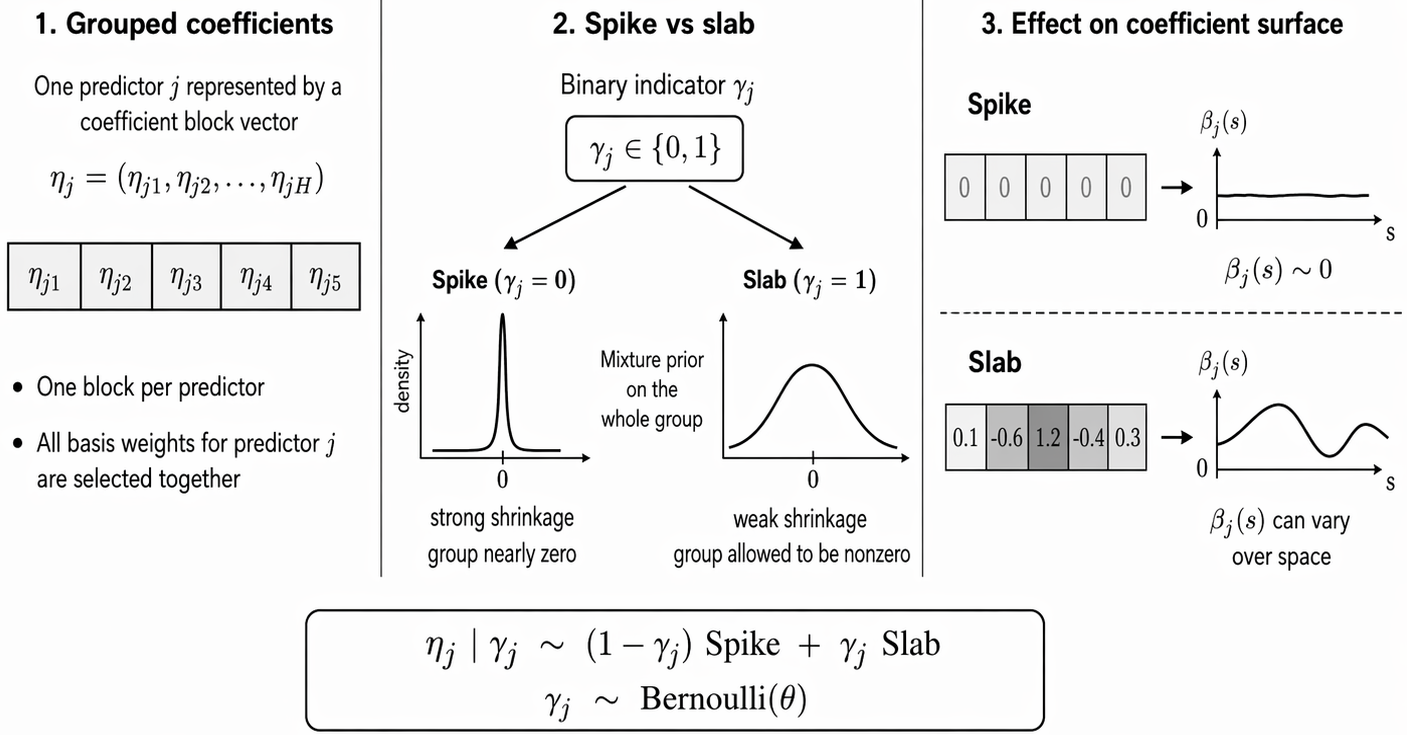
SSGL-SVC: Prior Setting
- Spike-and-Slab Group Lasso (Multivariate Laplace)
p(\boldsymbol{\eta}\mid \boldsymbol{\gamma}) = \prod_{j=1}^{p}\Big[(1-\gamma_j)\,\Psi(\boldsymbol{\eta}_j\mid \lambda_0) + \gamma_j\,\Psi(\boldsymbol{\eta}_j\mid \lambda_1)\Big], \quad \lambda_0 > \lambda_1 > 0
\Psi(\boldsymbol{\eta}_j\mid \lambda) \propto \exp\big(-\lambda \lVert \boldsymbol{\eta}_j \rVert_2\big)
\eta_j \mid \tau_j,\sigma^2 \sim N(\bm{0},\sigma^2\tau_j\hbox{\bf I}_H), \qquad \tau_j\mid \gamma_j \sim Ga\!\left(\frac{H+1}{2},\frac{\lambda_{\gamma_j}^2}{2}\right),\qquad \lambda_{\gamma_j}= \begin{cases} \lambda_0,&\gamma_j=0,\\ \lambda_1,&\gamma_j=1. \end{cases}
- The posterior inclusion probability (PIP) P(\gamma_j=1\mid\mathbf y) summarizes the posterior probability that predictor x_j has a non-negligible (spatially varying allowed) coefficient surface.
- Other priors \sigma^2\sim IG(a_\sigma,b_\sigma), \quad \gamma_j\mid \theta \sim \mathrm{Bernoulli}(\theta), \quad \theta \sim \mathrm{Beta}(a,b)
Posterior Computation
A Gibbs sampler is developed.
-
Posterior draws
- \boldsymbol{\eta} from normal
- \sigma^2 from inverse gamma
- \boldsymbol{\tau} from generalized inverse Gaussian
- \boldsymbol{\gamma} from Bernoulli
- \theta from beta
Derived coefficient surfaces \beta_j(s)=\mathbf{B}(s)^\top \boldsymbol{\eta}_j
Low-rank basis expansion makes each update depend on pH, not pn, spatial coefficients.
Low-rank spatial representations are a common strategy for scaling spatial models (Banerjee et al. 2008; Guhaniyogi et al. 2022).
Simulation Design
| Component | Design |
|---|---|
| Sample size | n=1000, 2000, 10000 |
| Predictors | p=5,7,10 |
| Metrics | prediction error, surface MSE, selection accuracy, time |
-
Basis-based
- Gaussian-SVC: No shrinkage, and ridge-type prior on each basis coefficient
- BSGL-SVC (Continuous Bayesian spatial group lasso) (Zhan et al. 2026): Continuous shrinkage with no spike-and-slab
- GGP-GAM (Geographical Gaussian process generalized additive model) (Comber, Harris, and Brunsdon 2024): frequentist equivalent of BSGL-SVC
-
Nonbasis-based
- MGWR (Multiscale geographically weighted regression) (A. Stewart Fotheringham, Yang, and Kang 2017): non-basis frequentist benchmark
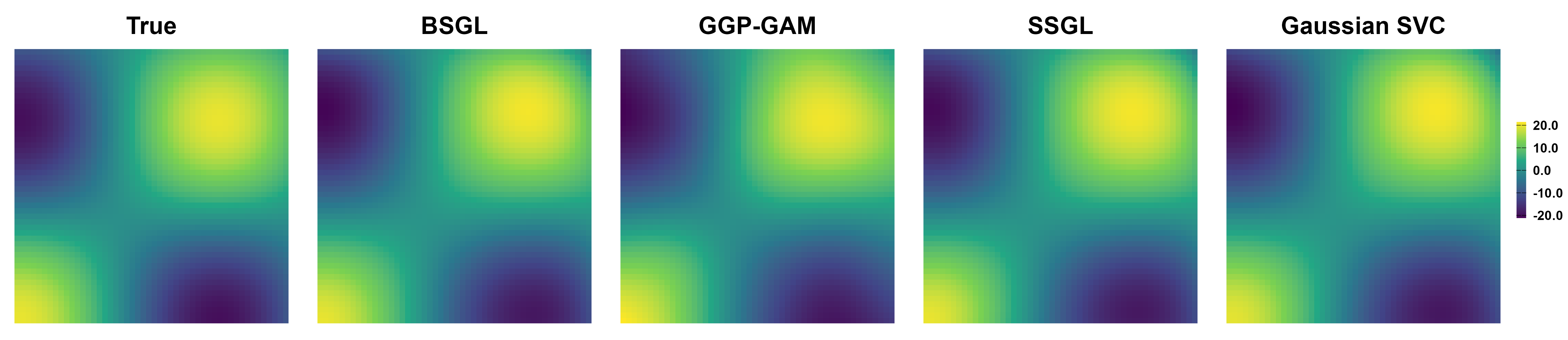
Simulation Results: Surface Recovery
All competing methods recover the main spatial patterns of active predictors with large signals
Simulation Results: Variable Selection
SSGL-SVC separates signal predictors from noise predictors more sharply than continuous shrinkage alone, BSGL-SVC and Gaussian-SVC.
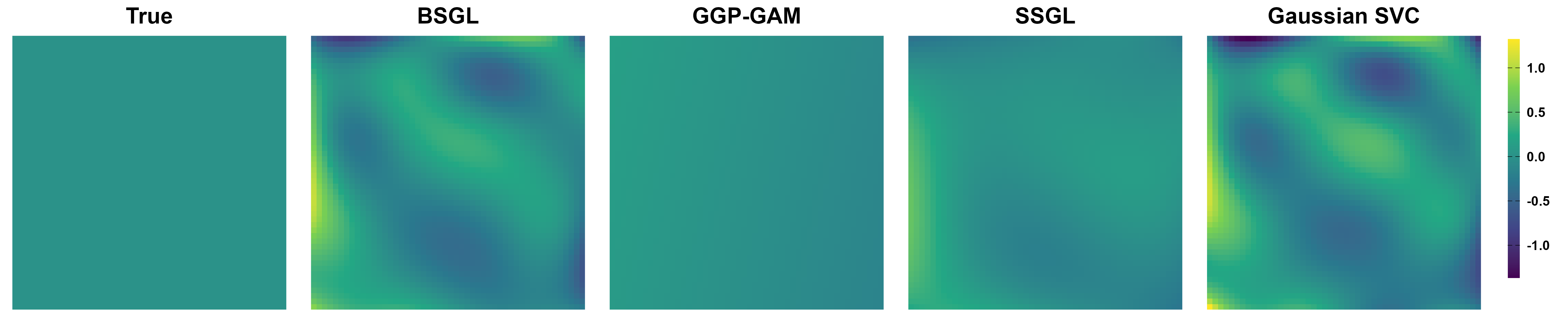
- GGP-GAM seems to work well, but . . .
Simulation Results: Unreliable Spatial Patterns
- GGP-GAM and MGWR produce spurious spatial patterns.
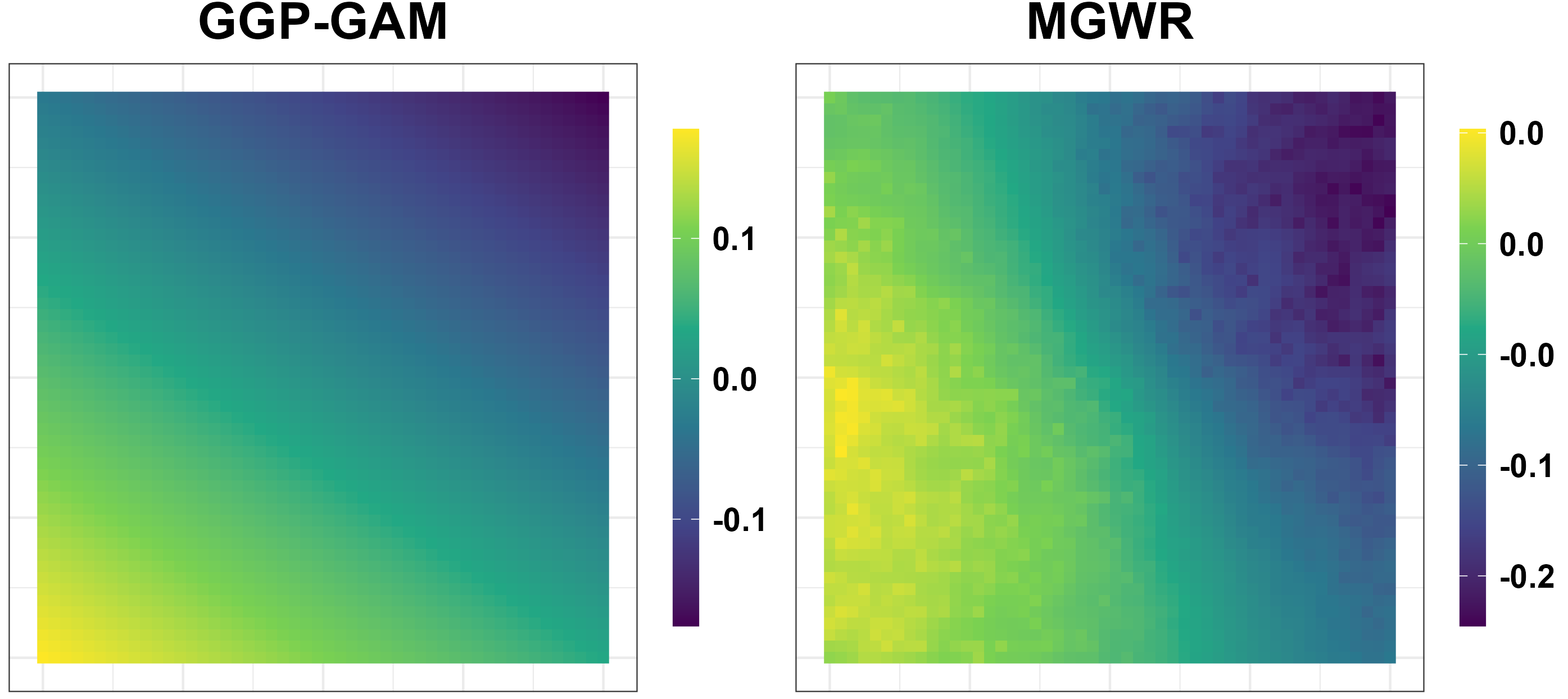
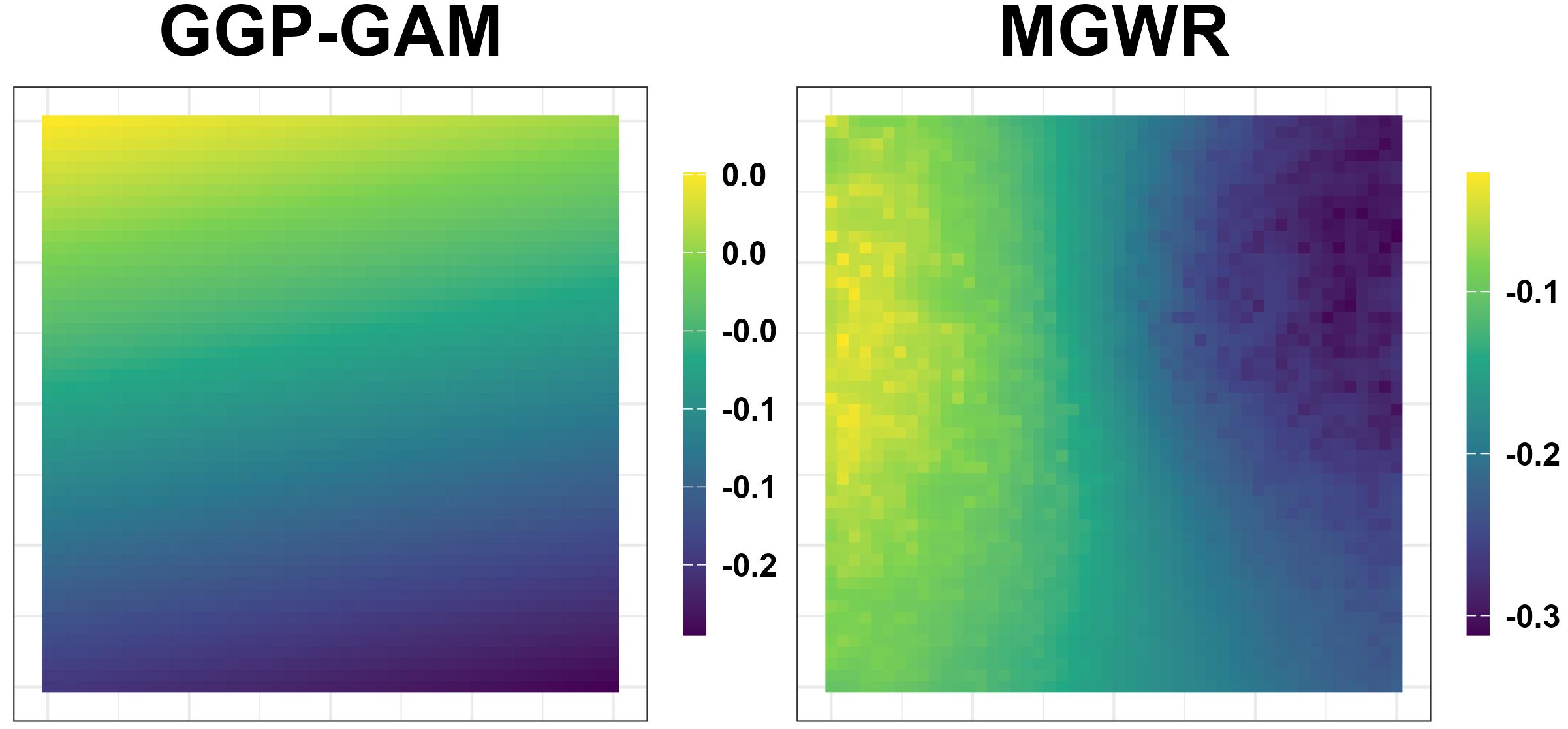
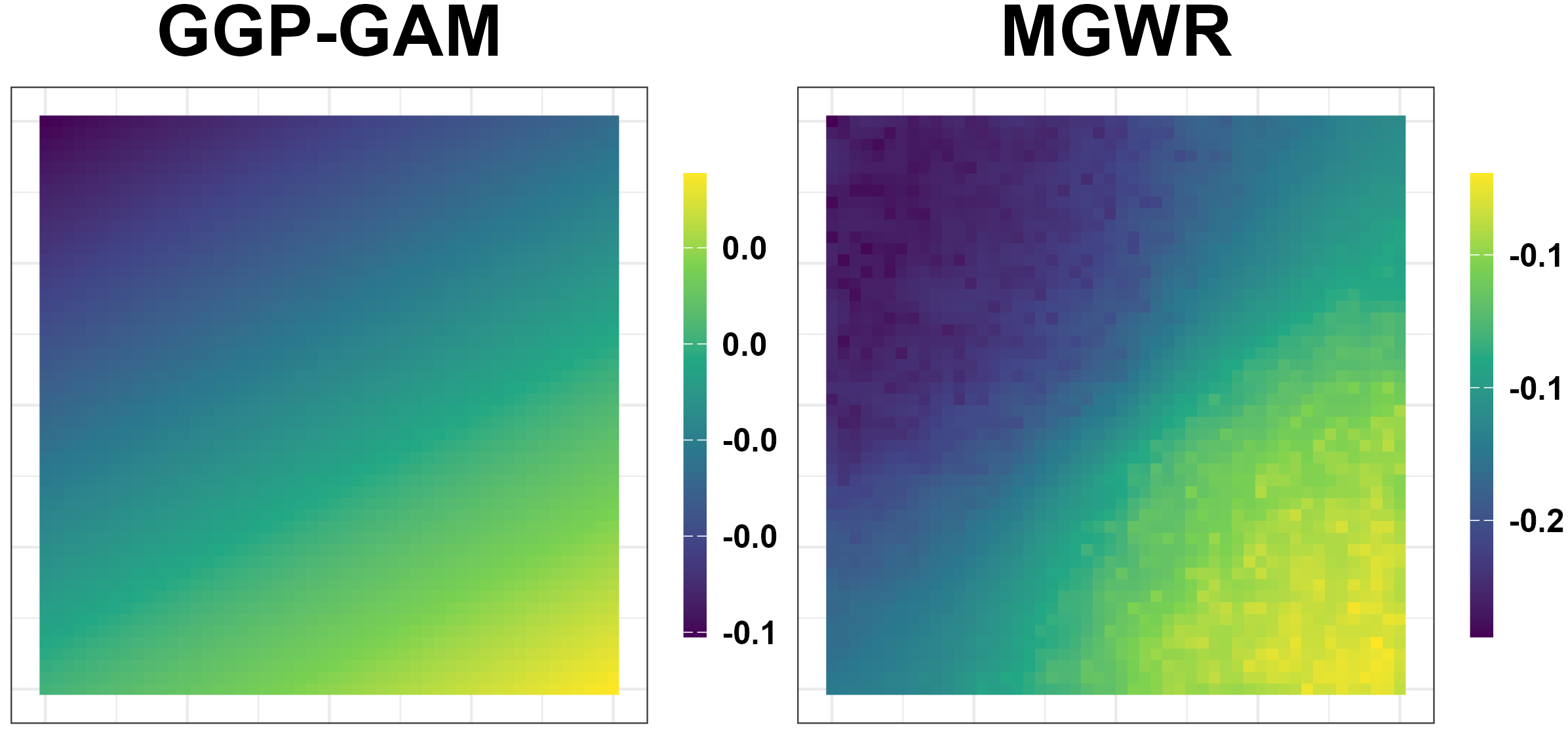
Simulation Results: Performance Measures
SSGL-SVC is competitive in prediction and improves shrinkage of inactive surfaces, especially at smaller n.
| Method | MSPE | MSE_1 | MSE_0 | MSE_{\text{avg}} |
|---|---|---|---|---|
| n=1000 | ||||
| Gaussian-SVC | 1.18 | 0.78 | 0.85 | 0.81 |
| BSGL-SVC | 1.14 | 0.74 | 0.40 | 0.60 |
| GGP-GAM | 1.10 | 0.89 | 0.07 | 0.56 |
| SSGL-SVC | 1.10 | 0.68 | 0.12 | 0.46 |
| n=10000 | ||||
| Gaussian-SVC | 1.03 | 0.22 | 0.18 | 0.20 |
| BSGL-SVC | 1.03 | 0.20 | 0.13 | 0.17 |
| GGP-GAM | 1.02 | 0.15 | 0.04 | 0.10 |
| SSGL-SVC | 1.01 | 0.18 | 0.05 | 0.12 |
Real Data Application
- MODIS (Moderate Resolution Imaging Spectroradiometer) by NASA on the region 30°N-40°N latitude, 104°W-130°W longitude
Response: Enhanced Vegetation Index (EVI) measuring vegetation greenness (Huete et al. 2002)
-
10 predictors:
- Spectral Reflectance: Red, NIR, Blue, MIR
- Ecosystem: GPP (amount of carbon through photosynthesis), LE (heat flux)
- Satellite Observation Geometry: View/Sun zenith angle, Azimuth angle
- Land Surface: LC Type4
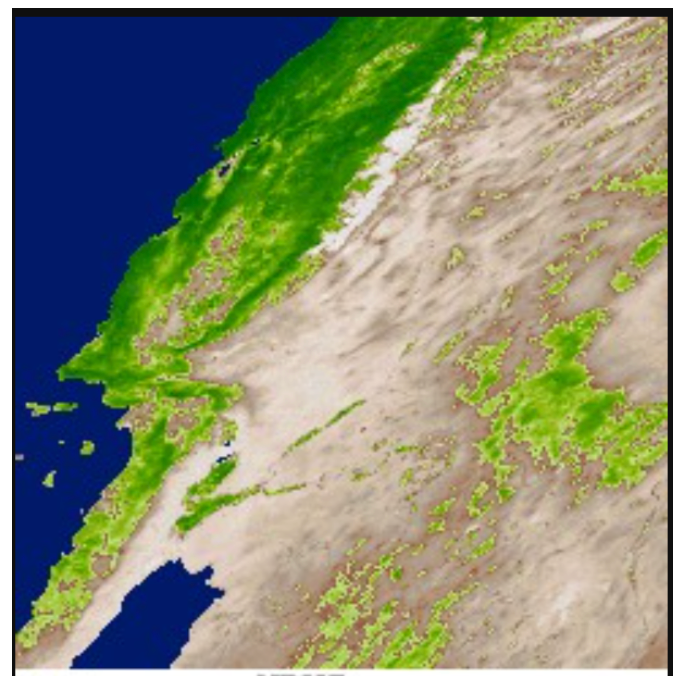
Which predictors have spatially varying associations with EVI?
MODIS: Coefficient Surface and Selection
SSGL-SVC leads to larger coefficient magnitudes.
\text{PIP} \approx 1 shows Red Reflectance is associated with EVI, and it has a non-negligible coefficient surface.
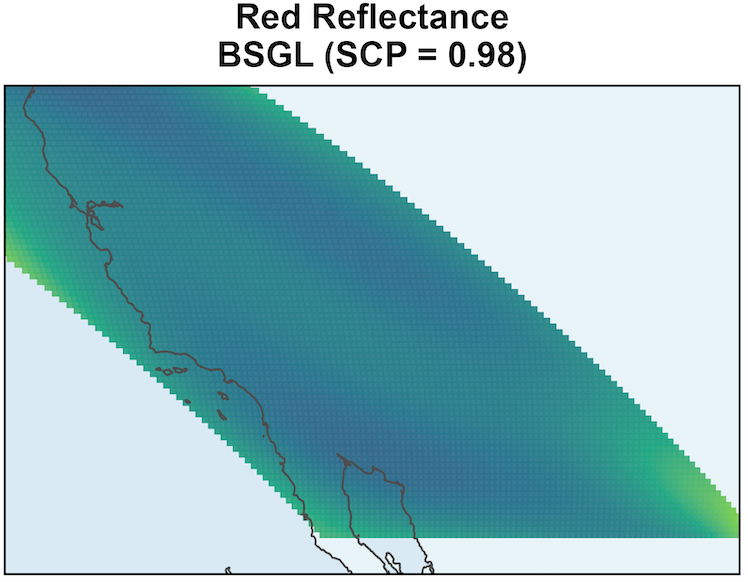
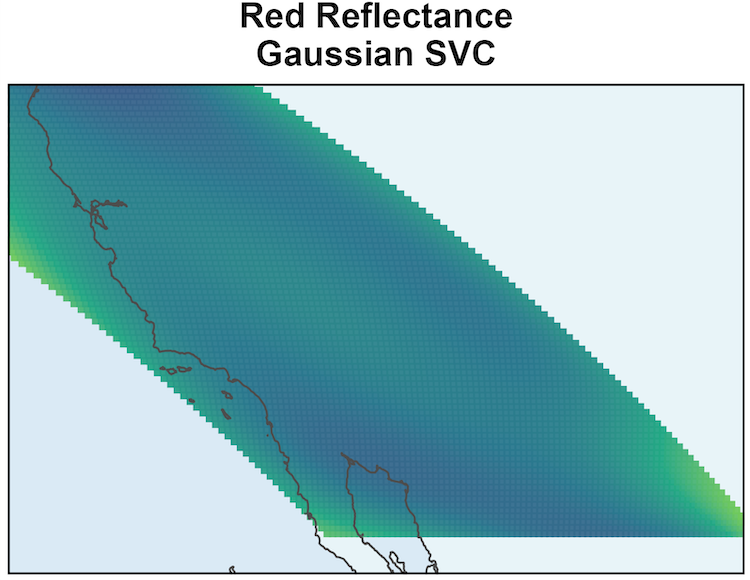
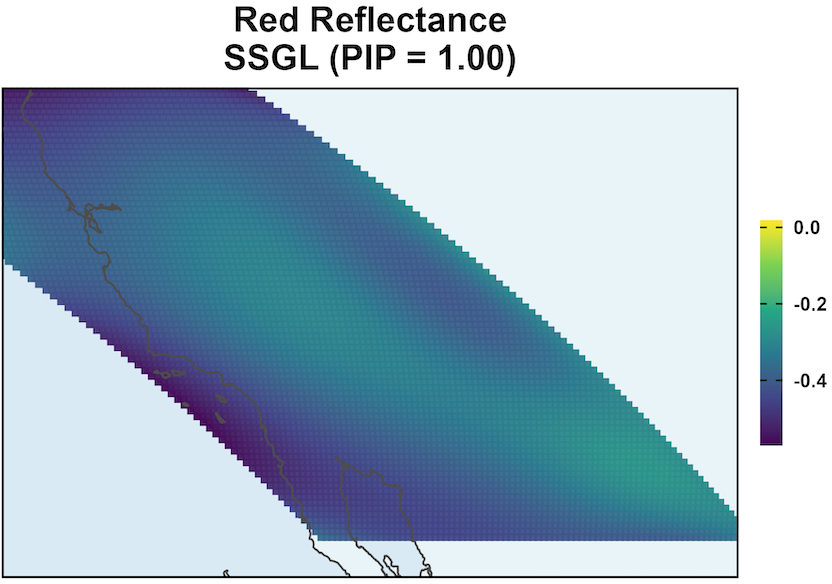
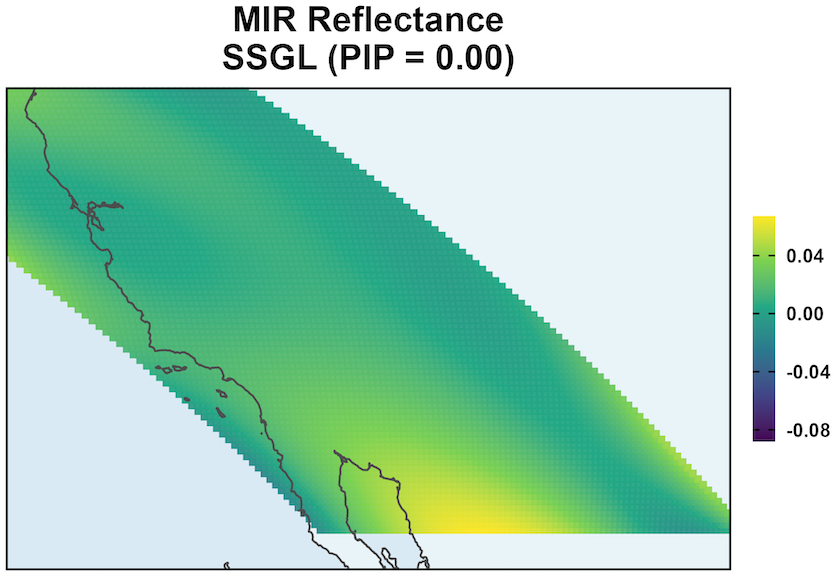
MODIS: Coefficient Surface and Selection
SSGL-SVC shrinks coefficients more toward to 0 when the effect is negligible.
\text{PIP} \approx 0 shows that given all other variables, MIR Reflectance’s effect is negligible.
Remaining variation in \beta_{MIR}(s) is due to residual posterior uncertainty.
The surface should not be interpreted as meaningful spatial signal.
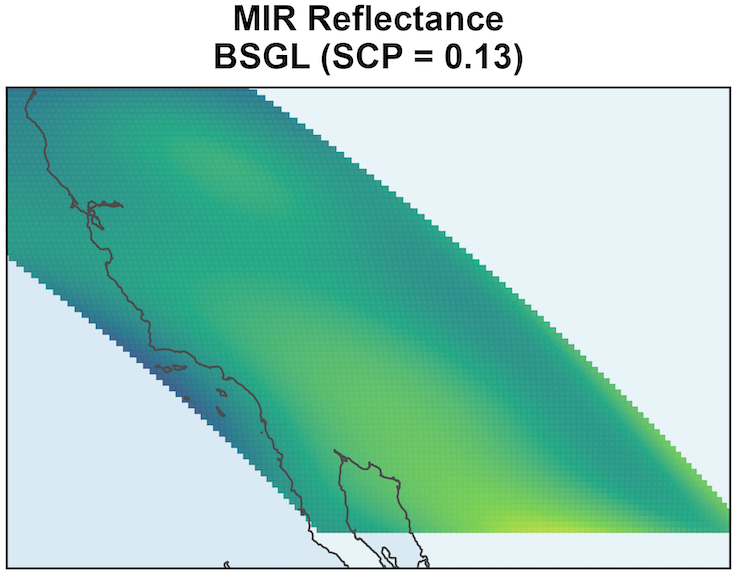
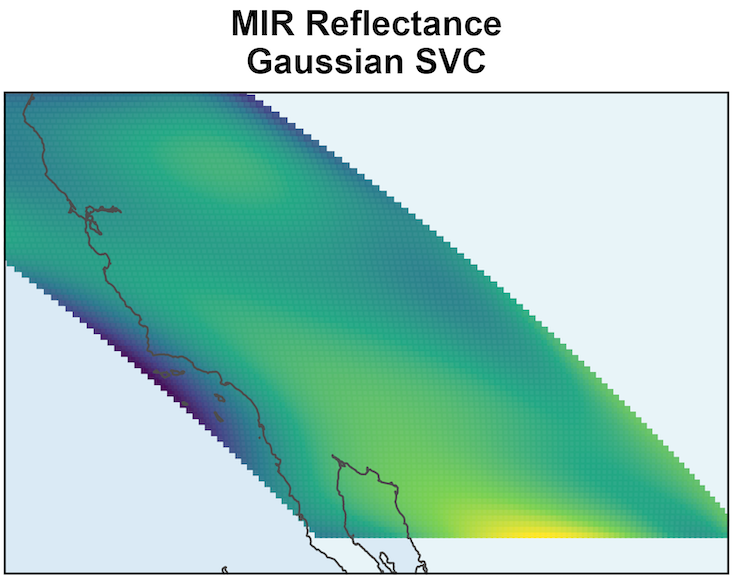
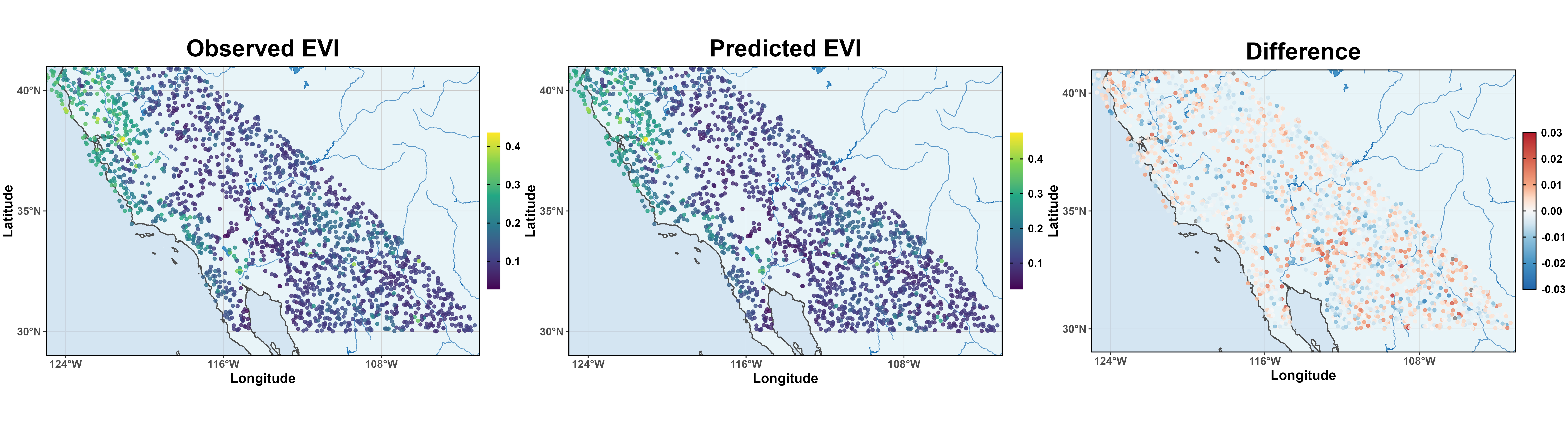
MODIS: EVI Prediction
Moran’s I verified that there is no spatial correlations in posterior residuals.
Great MSPE 8.38 \times 10^{-5} and better than BSGL-SVC and Gaussian-SVC.
Take Home Messages
- We propose a novel SVC model
- scalable via basis function approximation
- variable selection via spike-and-slab group lasso
- uncertainty quantification via Bayesian posterior probabilities
- Work in process and future work includes
- spatiotemporal modeling: AR noises, \beta(s, t), \gamma_j(t), etc
- Non-Gaussian responses: binary, counts, zero-inflated counts, etc
- irregular or constrained space: smooth within region, abrupt across boundaries, within-surface local sparsity selection
- scalable online learning: variational, EM + Laplace, NNGP / Vecchia / SPDE / sketching, etc